This paper moves beyond simply measuring "lifespan" as a single number. Instead, it maps genetic effects using age-dependent mortality (hazard) as the metric, identifying 29 lifespan loci (Vita) and 30 body-mass–lifespan trade-off loci (Soma). Crucially, most genetic effects and gene–gene interactions (epistasis) manifest as completely distinct networks by sex, meaning that sex-pooled analyses can miss or distort important signals. Several loci are further narrowed to candidate genes and connected to C. elegans experiments and human genetic data (Mendelian randomization), offering threads that may translate into human healthspan research.
1. Why "Lifespan Alone" Was Not Enough: Framing the Problem 🧩
The authors begin by noting that we still cannot fully explain why aging rate, lifespan, and overall mortality differ so much among individuals in humans, mice, and other organisms. Many variants linked to age-related disease risk have been found, but most of those may reflect the proximate causes of death (diseases, etc.) rather than the rate of aging itself.
"We do not understand the genetic, molecular, cellular, or organismal processes that create variation in the rate of aging, lifespan, and overall mortality in humans, mice, or other model organisms."
The researchers therefore set out to "separate cause from consequence" and design a new approach to find variants that alter mortality across the entire lifespan—from early life through old age. The paper addresses four main questions:
- Which DNA variants affect mortality and lifespan, and when do they act? (durable / transient / late-life)
- How do sex differences in mortality (including pre- and post-reproductive periods) map to specific loci? (G×S, G×G, antagonism between sexes)
- Which loci create the opposing correlation where large early body size predicts shorter life but large late-body size predicts longer life?
- How do these dynamics fit the major evolutionary theories of aging (mutation accumulation, antagonistic pleiotropy, somatic maintenance cost / disposable soma)?
2. The Stage: Large-Scale ITP Mouse Data and "Survivorship" Mapping 📈
The researchers use data from the U.S. NIA Interventions Testing Program (ITP), drawing on UM-HET3 mice—a genetically diverse, outbred population accumulated over more than 20 years across three testing sites. After genetic quality filtering, 6,438 animals are included in the analysis.
- Youngest survivorship cohort: T42 (survived to at least 42 days)
- Oldest survivorship cohort: T1,100 (559 animals that survived past 1,100 days)
- First observed death: day 46; last observed death: day 1,456
The core technical innovation is actuarial mapping: repeatedly mapping lifespan/mortality loci while progressively left-truncating the cohort (i.e., restricting analysis to animals that survived past successively older ages—T42, T365, T740, etc.). This tracks when each genetic effect strengthens or weakens along the time axis.
Compared to a prior study (3,055 mice, 7 loci), this work doubles the sample and quadruples the markers while applying more powerful actuarial mapping, yielding:
- ~4× more Vita loci → 29 total
- 30 new Soma loci linking body mass to lifespan
- Systematic characterization of age- and sex-varying heritability and sex-specific epistasis
3. Result 1: 29 Vita Loci—Decomposing "When Mortality Risk Diverges" by Locus 🧬
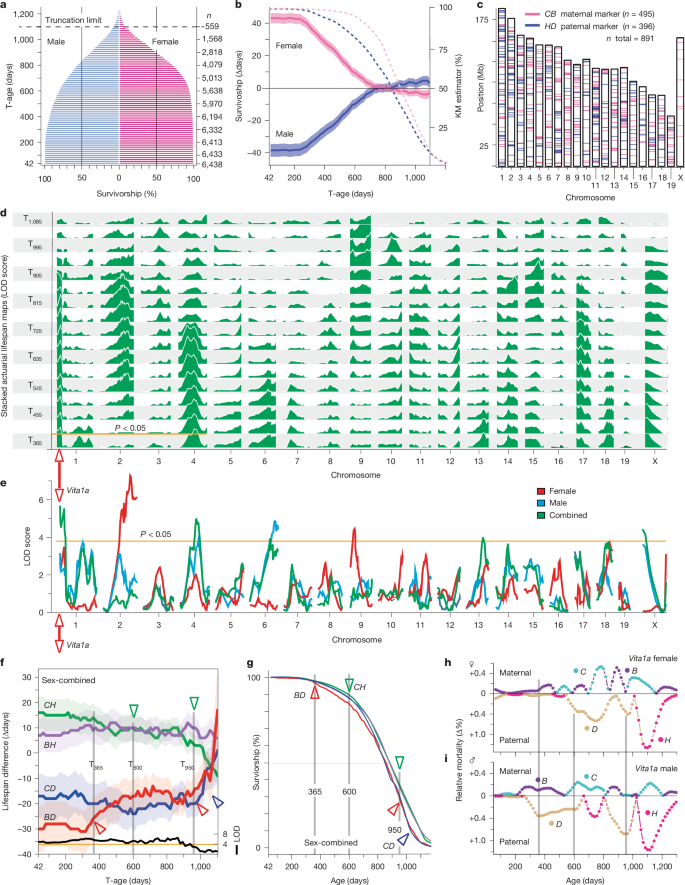
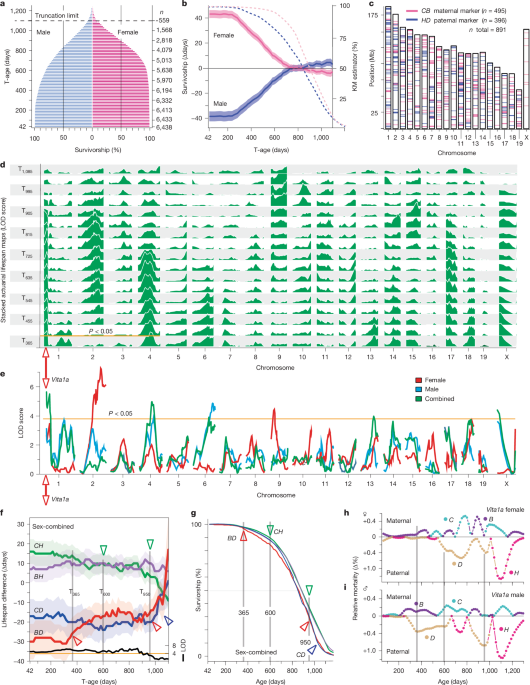
3.1 The Sex Gap in Lifespan Is Large Early, Then Disappears
In the T42 cohort, mean lifespan is 806 days for males and 887 days for females—an 81-day female advantage. Male deaths cluster heavily between days 215 and 410, narrowing the gap; by T740, expected lifespan converges at ~946 days for both sexes. In the oldest cohorts (T935–T1,100), males show a ~8-day advantage, possibly partly attributable to males being housed individually at older ages.
3.2 Vita Loci Average 36 Days of Effect, But Shape Varies by Age and Sex
Applying stringent multiple-testing corrections (FDR, Bonferroni, Cauchy combination tests), the researchers confirm 29 Vita loci.
- Mean locus effect: ~36 days
- Variance explained at peak: females ~2.5%, males ~3.2%
- Mean active age range per locus: ~349 days (varies widely across loci)
Notably, the same locus can show effect reversals—a genotype disadvantageous early becomes advantageous late, or vice versa.
"The effect of Vita loci is age-dependent."
4. Result 2: Dynamics of Vita Loci and Time-Varying Heritability ⏳
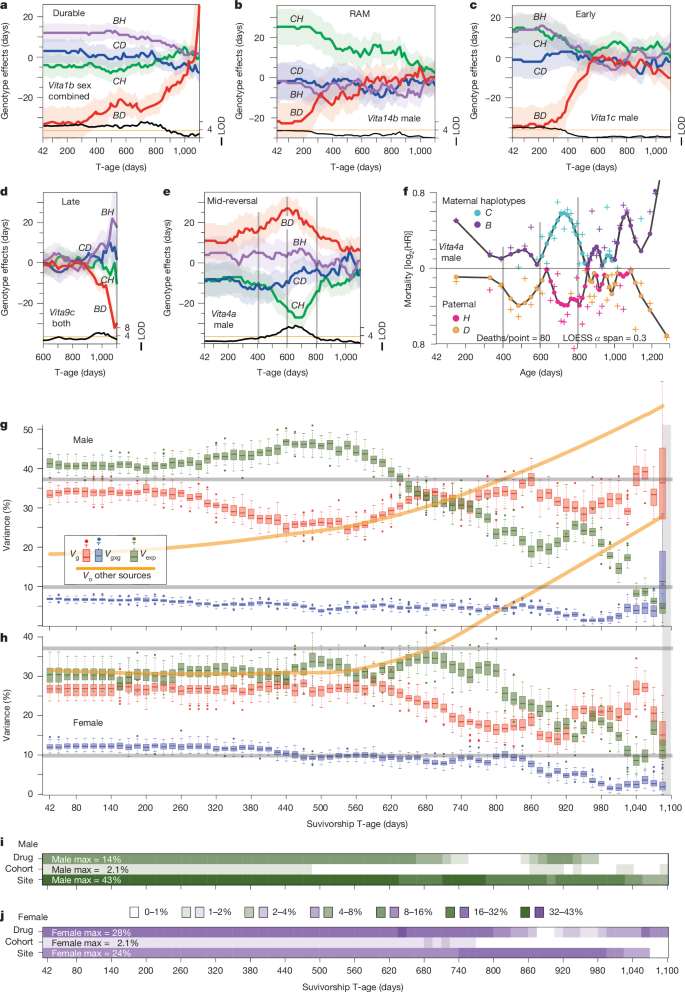
By examining actuarial effect curves (expected-lifespan differences across survivorship cohorts), Kaplan–Meier survival curves, and age-specific mortality differences (hazard ratios), the authors classify Vita loci into four broad types:
- Durable: Relatively consistent predictive power from early to late life (e.g., Vita1a, Vita1b)
- "Consistent" does not mean the mortality effect is age-independent; some of these loci concentrate their mortality difference in late life.
- Steadily diminishing: Large early difference that converges with age
- Includes types that superficially resemble candidates for regulating aging rate (RAM)
- Age-range restricted, further split into:
- Early (roughly before day 500), mid-life (days 500–845), late-life (after day 860)
- Early-acting loci are ~3× more common in males
- Reversal: Effect direction flips with age (12 loci)
Heritability "Fluctuates More in Males, Stays Flatter in Females"; Epistatic Variance Is Larger in Females
- The genetic variance (Vg) explained by the 29 Vita loci is higher in males early on (T42–T320; up to ~40%), drops around T500 during the early male mortality wave, then rises again in old-age cohorts (up to ~50%).
- Females hold a relatively stable ~27% through T620, then decline, with partial recovery in the oldest cohorts but remaining lower than males.
- In contrast, two-locus epistatic variance is ~2× higher in females than males across reproductive-age cohorts (12% vs. 6%).
Environmental factors—testing site, cohort year, and supplement group (individually non-significant but jointly significant)—account for substantial variance. The site effect alone reaches up to 43% in males and 24% in females.
5. Result 3: Sexual Antagonism—The Same Locus Can Benefit Females While Harming Males ⚖️
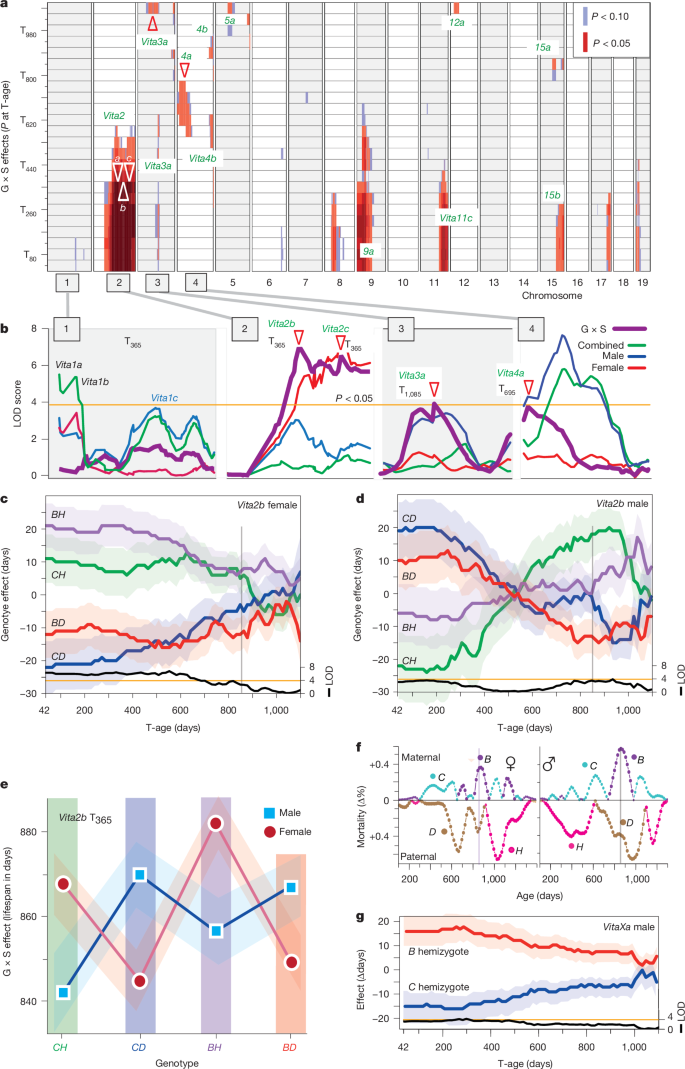
Strong genotype × sex (G×S) interactions are confirmed at 14 Vita loci. The clearest example is Vita2b on chromosome 2: the same genotype
- extends female expected lifespan through T700, but
- acts disadvantageously in males during certain age windows, and
- in males, early benefits reverse into late-life costs after day 600—a pattern consistent with antagonistic pleiotropy
The authors warn that sex-pooled mapping can generate spurious peaks (artifacts) by obscuring these interactions.
The male-specific VitaXa on the X chromosome spans a broad region (~70 Mb proximally), and because males are hemizygous, the signal may reflect the aggregate net effect of multiple recessive variants acting without a heterozygous counterpart.
6. Result 4: Soma Loci—Genetically Decomposing the "Large Body vs. Long Life" Trade-off 🐭⚙️

Body mass measured at days 42, 183, 365, 548, and 730 is correlated with subsequent lifespan, and these correlations are tracked actuarially as they shift with age and sex.
- Body mass at days 42–548 generally correlates negatively with subsequent lifespan (heavier → shorter-lived)
- The effect is largest around day 183 (~6 months, around reproductive maturity):
- Male correlation: rho ≈ −0.28
- Female correlation: rho ≈ −0.11
- Converted to "lifespan cost per gram of body mass" at the reproductive-age peak:
- Males: −14.3 days/g
- Females: −3.7 days/g
- At very late ages (day 730 body mass), the correlation turns positive in some windows, consistent with late-life body weight reflecting resilience.
A key insight is that loci governing body mass (Mass loci) and loci governing the body-mass–lifespan correlation (Soma loci) are largely non-overlapping. In other words, "genes that make you heavy" differ from "genes that determine how much being heavy costs you in lifespan."
Using correlation trait locus (CTL) mapping, the authors identify 30 Soma loci:
- 15 detected only in males, 4 detected only in females (strong male bias)
- 19 loci modulate the early (days 42–185) negative correlation
- 11 loci modulate the post-reproductive positive correlation
- Locus effect sizes: 2–29 days/g
Soma3b is strong in males but nearly absent in females; Soma11a is the strongest female Soma locus, though smaller in absolute magnitude than the most extreme male effects.
7. Result 5: Epistasis Networks Are Sex-Segregated 🕸️
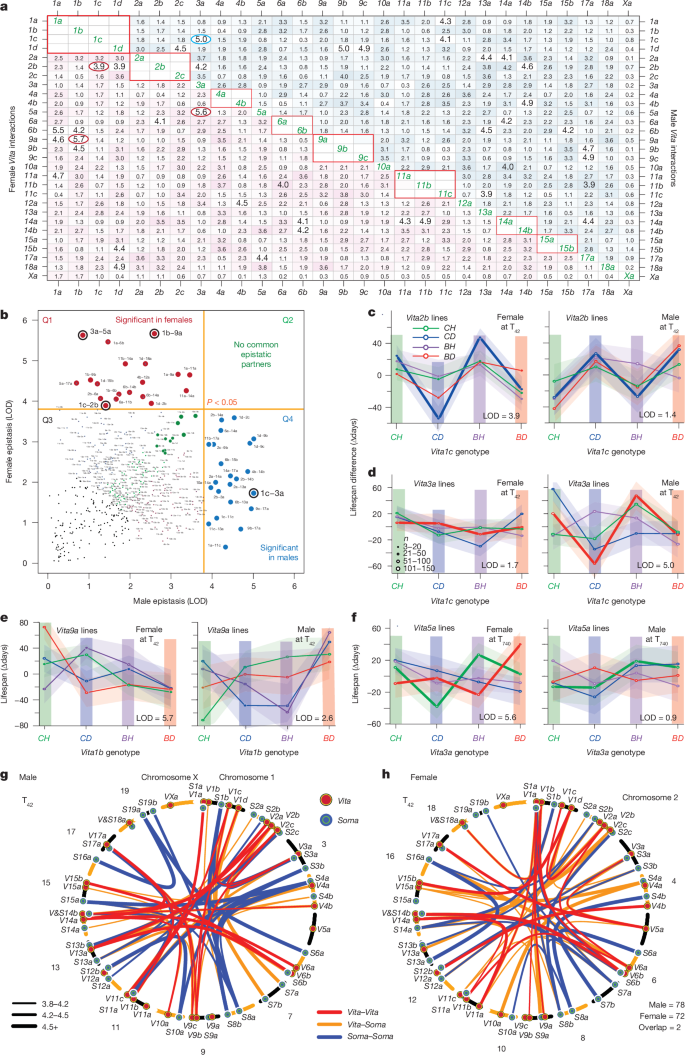
The researchers test all pairwise interactions among the 59 loci (Vita 29 + Soma 30) at four survivorship ages (T42, T365, T740, T905), separately by sex. The number of interactions is large, but the most striking finding is that female and male epistatic networks share almost no interactions.
- Significant Vita–Vita interactions at T42 (stringent correction): 22 in males, 19 in females
- Across all four survivorship cohorts combined: 59 in males, 43 in females (Vita–Vita)
- Interactions significant in both sexes simultaneously: virtually none (near-zero overlap)
The authors frame this finding as:
"These results are consistent with a strongly sex-dimorphic genetic architecture—which we speculate reflects a form of 'genetic diplomacy' that reconciles distinct life-history strategies between the sexes."
Female epistatic partnerships are relatively stable across age cohorts, while male interaction pairs are more substantially reshuffled as cohorts age.
8. From Map to Mechanism: Candidate Genes, Worm Experiments, and Human Data 🔬
To move from locus maps toward actual mechanisms, the researchers focus on more tractable loci and chain together candidate gene prioritization → model-organism experiments → human genetic data.
8.1 Vita1a: A Small Interval (14 Genes) Highlights Autophagy, Mitochondria, and Nutrient-Sensing Candidates
Vita1a overlaps only 14 protein-coding genes—a compact locus. Highlighted candidates include:
- Mrpl15 (mitochondria-related)
- Atp6v1h (lysosomal acidification, linked to mTOR function)
- Rb1cc1 (nutrient sensing / autophagy)
Atp6v1h is noted for prior evidence linking it to lifespan extension in flies and worms and to metabolic disease risk in humans. The locus also shows nonlinear interactions with specific other loci exclusively in females, flagged as an important constraint when validating candidate genes.
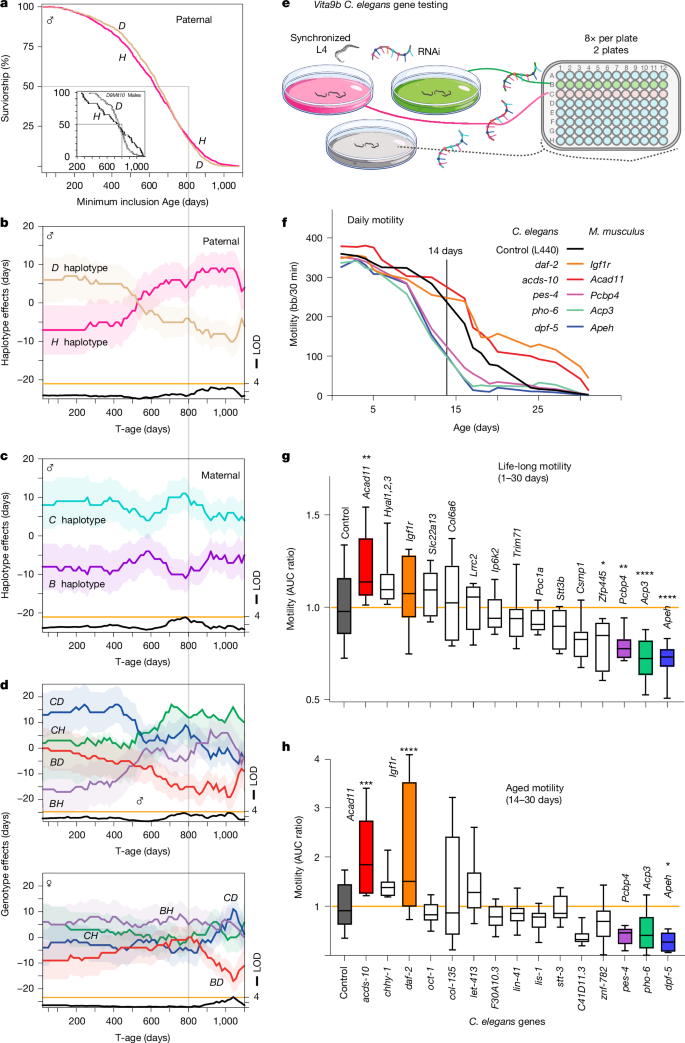
8.2 Vita9b: Replicated Across Historical Cohorts + Motility Changes in C. elegans RNAi
Vita9b affects both sexes in old survivorship cohorts and has been observed in earlier independent cohorts—a replicated locus. Candidate genes in this interval were knocked down by RNAi in worms, and motility (a proxy for healthspan) was measured.
Knockdown of acds-10 (ortholog of mouse Acad11) improved late-life motility (after day 14) in a pattern resembling knockdown of the longevity-positive control daf-2 (ortholog of mouse Igf1r). Knockdown of several other candidates had the opposite effect, reducing motility.
8.3 In Human Data (Mendelian Randomization), APEH Stands Out
Among mouse Vita9b candidates tested in human MR analysis using expression quantitative trait loci (eQTLs) as instruments, APEH shows:
- Significant association with parental lifespan (especially paternal) and personal longevity measures
- Higher APEH expression in blood positively associated with male longevity
By contrast, candidates testable from Vita1a (LYPLA1, MRPL15, PCMTD1, TCEA1) yielded no clear MR signals. The paper thus demonstrates that some loci translate cleanly to human data while others remain unresolved.
9. Discussion: Testing Three Evolutionary Theories of Aging with Age-Specific Genetic Effects 🧠
Where prior work treated lifespan as a single number, this study tracks age-specific mortality changes from puberty through senescence by genotype and sex, showing how 59 loci sculpt the mortality curve.
- Mutation accumulation theory: Variants harmful only after reproduction face relaxed selection and accumulate → late-acting Vita loci are consistent with this view
- Disposable soma theory: Energy investment in reproduction and growth trades off against somatic maintenance → Soma loci that amplify the early negative body-mass–lifespan correlation exemplify a genetic implementation of the investment-vs-maintenance trade-off
- Antagonistic pleiotropy: Early benefit (survival/reproduction) ↔ late cost (increased mortality) → Vita2b, Vita9b, and others whose effects reverse with age are the clearest examples
For the early male mortality excess, the authors raise male–male competition, aggression, and stress as possible contributors while acknowledging that a complete pathological explanation remains outstanding. They also recall that ITP drug interventions generally work better in males (with rapamycin as an exception), and suggest the loci defined here could serve as anchors for testing drug–locus interactions in future work.
10. A Key Interpretive Point from Methods: "Starting Age (Left Truncation) Changes the Mean Lifespan" 📌
One striking methodological note is that reported mean lifespan depends critically on which ages are included—not just on birth. The study's baseline inclusion criterion is day 42 (around puberty, when tail tissue is collected), so all lifespan figures in the paper are conditioned on survival to that age.
"Researchers often incorrectly imply that mean lifespan is calculated from birth, ignoring earlier (often unknown) deaths. … We can only measure lifespan for UM-HET3 offspring that survived to puberty."
This applies equally to human studies: how a longevity cohort is defined has large consequences for interpreting results.
11. Data and Code Availability: A Foundation for Replication and Extension 🧾
Primary data are provided in supplementary tables, with additional data, tools, and figures released publicly:
- Full lifespan/body-mass data (with tools): https://genenetwork.org/show_trait?trait_id=10001&dataset=HET3-ITPPublish
- Figures and supplementary portal: https://aging.genenetwork.org/UM-HET3 https://files.genenetwork.org/current/umhet3_2025/
- Code (GitHub): https://github.com/DannyArends/UM-HET3
12. Conclusion
This study compellingly demonstrates—at scale—that aging genetics must be studied along the full time axis of life, and with sexes analyzed separately. The 29 Vita and 30 Soma loci break down the outcome of "lifespan" into "when and how mortality risk rises," and the finding that epistatic networks are almost entirely sex-segregated is a direct warning for how future aging and intervention studies should be designed. Beyond mapping, the Vita9b case shows that the chain from locus map → model-organism experiment → human genetic data can actually work—one of the paper's most consequential contributions.